Laboratory Trajectories Improve Kidney Failure Risk Estimation
Current kidney failure risk equations rely on only a patient's most-recent laboratory values, even though patients
often have years of routinely collected lab history. This project examines whether longitudinal lab
trajectories can improve risk estimation while remaining interpretable and clinically grounded.
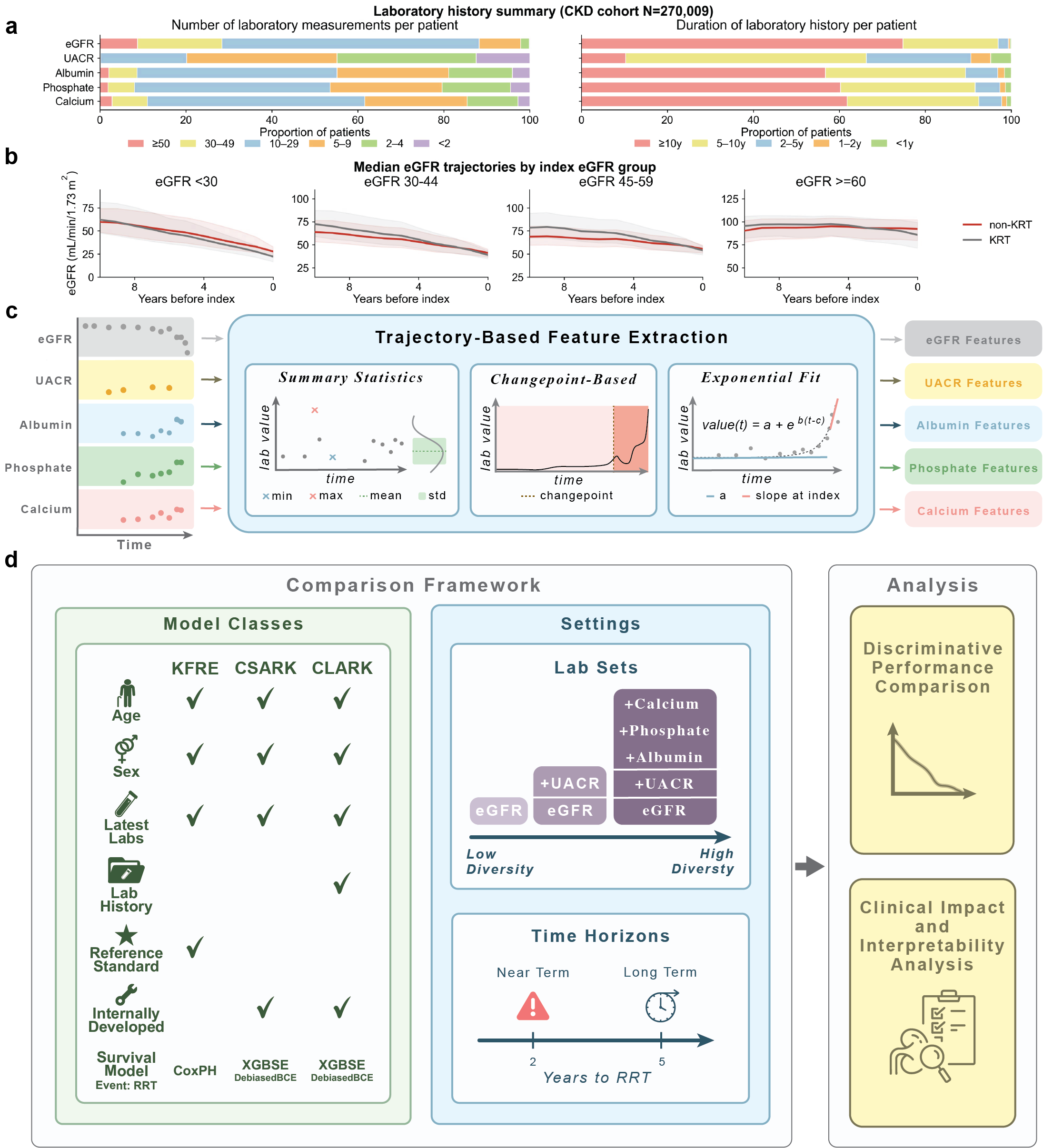
From a large EHR dataset spanning over 5 million individuals, we constructed a cohort of 270K patients with chronic
kidney disease. We developed a trajectory-based extension of standard risk models that incorporates features derived
from repeated laboratory measurements over time. These models improve identification of high-risk patients compared with
latest-value approaches, particularly for longer-term prediction.
The results highlight how routinely collected laboratory histories contain meaningful signal beyond a single measurement,
and how simple, interpretable trajectory features can improve risk stratification in clinical practice.


 Python (primary)
Python (primary) R
R Git
Git Docker
Docker